'Omics-related Data
Guidance for submitting genetic and other types of 'omics-related data
Contributing Genetic Accessions
We recommend that sequence data be submitted to national biological archiving services such as the National Center for Biotechnology Information (NCBI) GenBank, or the European Nucleotide Archive (ENA) where they can be unified under a BioProject in NCBI or Study in ENA. Related metadata and supporting environmental data should be submitted to BCO-DMO to further enable discovery and re-use (See Advantages of Sharing 'Omic Data Links through BCO-DMO).
What to send to BCO-DMO
If you have metadata and additional data related to your genetic accessions (collection date, latitude, longitude, taxon names, treatment descriptions, environmental measurements, etc.), please submit them to us along with any relevant accession numbers (e.g. BioSample, SRA run id, etc). Note that submitting your accession numbers and related data in a tabular format (e.g. Excel, comma- or tab-delimited) will expedite our processing of your data. We will then create a Dataset Landing Page at BCO-DMO which is linked to your Project Page.
To submit accessions and related data, use our online Submission Tool at https://submit.bco-dmo.org/ to enter your metadata and upload data files. See the "Submitting Data" page for more information on using the Submission Tool.
(If you cannot use the online submission tool, you can complete the DATASET.rtf metadata form and send it to info@bco-dmo.org along with any relevant data files.)
If applicable, please include the following information with your submission to BCO-DMO:
BioProject number
taxonomic names
description of the types of sequences
locations where species were collected (including latitude and longitude and cruise ID numbers, if known/applicable)
sequencing and analysis methods (including instrument names and models)
any other relevant information that will enable others to understand and re-use the data (e.g. published papers)
Example Datasets:
ezRAD libraries of Pocillopora spp. collected in 2019: https://www.bco-dmo.org/dataset/881853
Accession numbers for metagenome-assembled genomes (MAGs): https://www.bco-dmo.org/dataset/868323
Hurricane Harvey Coral Gene Expression: https://www.bco-dmo.org/dataset/817298
On these dataset landing pages, click the "View Table" or "Get Data" button at the top of the page to see the data published by BCO-DMO. Note that the related NCBI pages (like BioProject) are listed in a "Related Datasets" section on the landing page.
If you have any questions about how to submit data, please contact us at info@bco-dmo.org.
Advantages of Sharing 'Omic Data Links through BCO-DMO
'Omics is a term that refers collectively to a group of rapidly evolving multi-disciplinary fields (e.g. genomic, transcriptomic, proteomics, and metabolomic), each seeking to quantify and describe an entire collection of biological molecules of a particular type (e.g., genome). Ocean ecosystem investigators are increasingly using 'omics tools in their research due to their ability to document the environment's status in time and space. There are many benefits gained by submitting 'omics data links to BCO-DMO.
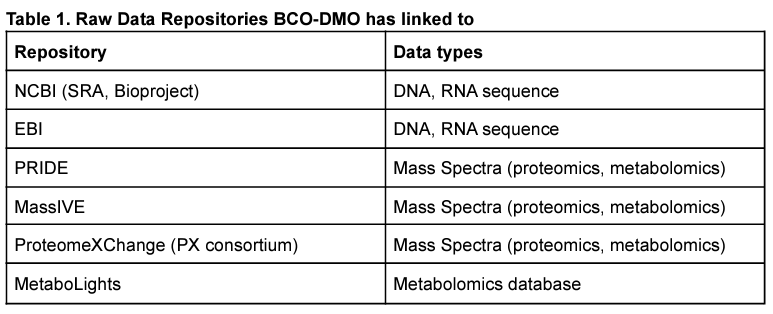
While there are dedicated repositories for 'omics data (Table 1), limited discoverability and accessibility of metadata and environmental data are major obstacles to reusing 'omic datasets. This is generally because the specialized 'omics repositories have been designed primarily for biomedical research and they often lack the ability to connect to environmental research or associated metadata. Although BCO-DMO does not host raw sequence or mass spectrometry data, we can easily link to the repositories that do. In doing so, BCO-DMO can allow other researchers to discover your data and place it in its appropriate environmental context. Because BCO-DMO datasets are connected to their expeditions, environmental data associated with the meta'omic sample locations can be easily connected. Similarly, BCO-DMO connects data to grants and/or projects, allowing laboratory-based experimental data to be associated with 'omic data. BCO-DMO's website is optimized for data discovery using search engines, and data types can be searched for within BCO-DMO's holdings. With a minor effort in submission, all the hard work put into collecting your data can be used to help other researchers throughout the world better discover and interpret your data. The data generators benefit from increased citations and collaborations.
Last updated